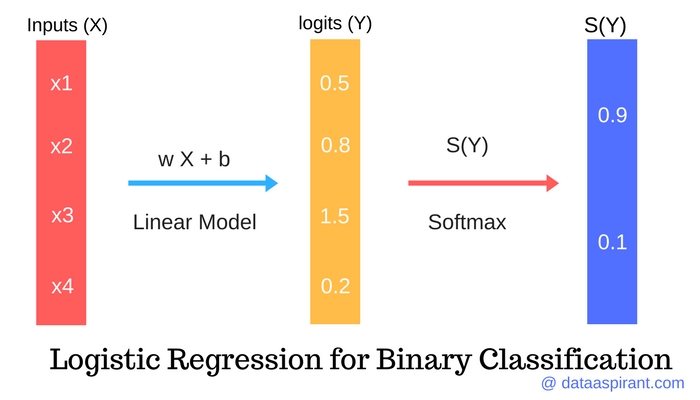
 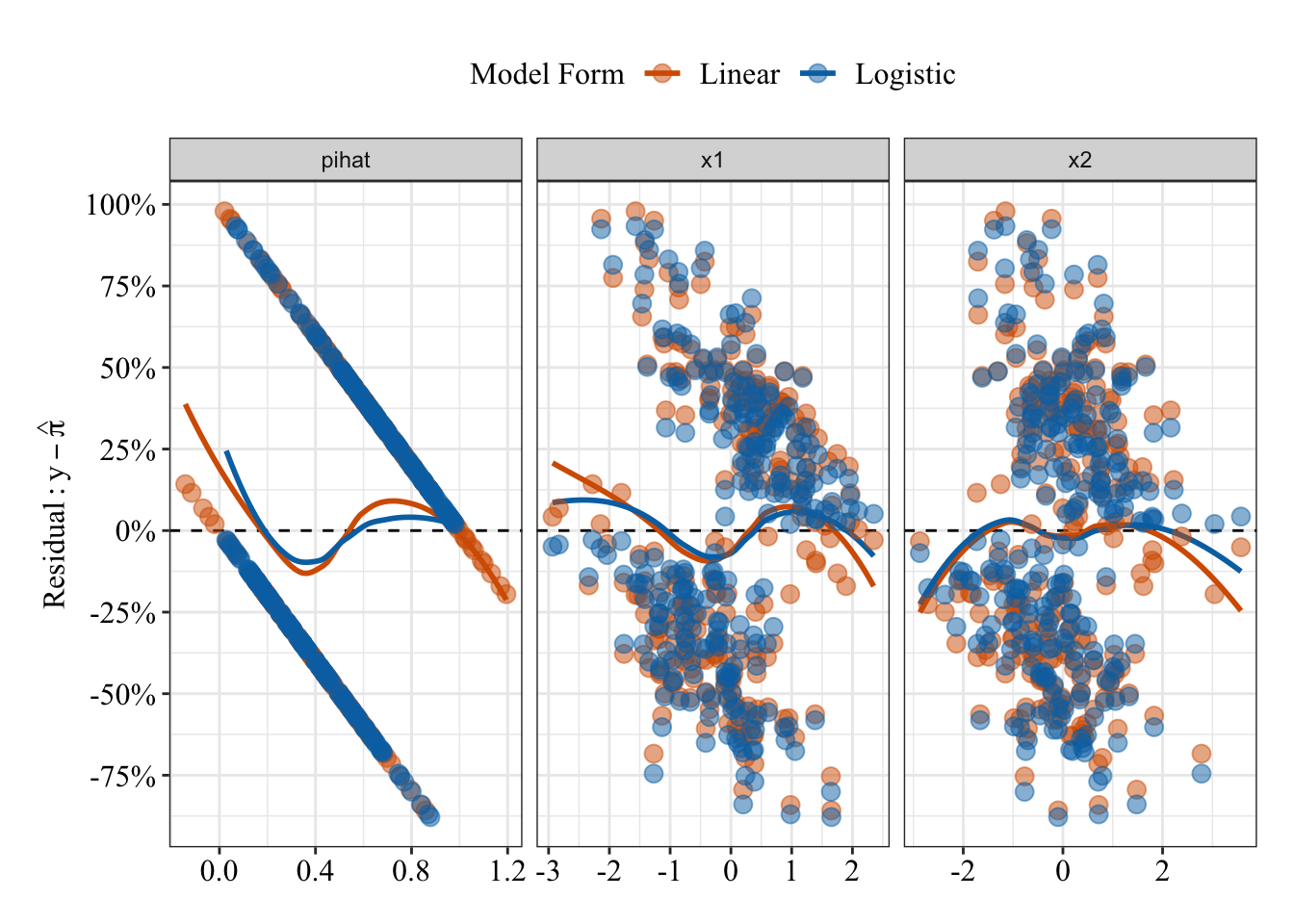 
Below I am going to describe how we do that mapping. Now the question arises, how do we map binary information of 1s and 0s to regression model which uses continuous variables? The reason we do that mapping is because we want our model to be capable of finding the probability of desired outcome being true. Note that the number 0 on Y-axis represents that half of the counts of total number is on left and half of total count is on right, but it cannot be the case always. If you have a greater number of 1s then that S will be skewed upwards and if you have greater numbers of 0s then it will be skewed downwards. To do that you have to imagine that the probability can only be between 0 and 1 and when you try to fit a line to those points, it cannot be a straight line but rather a S-shape curve. But how can we use that probability to make a kind of smooth distribution that fits a line (Not linear) as close as possible to all the points you have, given that those points are either 0 or 1. 
All we have is the counts of 0s and 1s which is only useful to find probabilities for example say if you have five 0s and fifteen 1s then getting 0 has probability of 0.25 and getting 1 has the probability of 0.75. Now, this is all good when the value of Y can be -∞ to + ∞, but if the value needs to be TRUE or FALSE, 0 or 1, YES or No then our variables does not follow normal distribution pattern. Also, the error term εi is assumed to be normally distributed and if that error term is added to each output of Y, then Y is also becoming normally distributed, which means that for each value of X we get Y and that Y is contributing to that normal distribution. We know this is linear because for each unit change in X, it will affect the Y by some magnitude β1. We have equation of the form Yi = β0 + β1X+ εi, where we predict the value of Y for some value of X. We know that a liner model assumes that response variable is normally distributed. Note that this is more of an introductory article, to dive deep into this topic you would have to learn many different aspects of data analytics and their implementations. When I was trying to understand the logistic regression myself, I wasn’t getting any comprehensive answers for it, but after doing thorough study of the topic, this post is what I came up with. Here I have tried to explain logistic regression with as easy explanation as it was possible for me. Ggplot(df, aes(d = known.I am assuming that the reader is familiar with Linear regression model and its functionality. # `m` (marker) holds the predictor values # the aesthetic names are not the most intuitive # combine linear predictor and known truth for training and test datasets into one data frameĭf <- rbind(ame(predictor = predict(, ),ĭata.frame(predictor = predict(, Pima.te), <- glm(type ~ npreg + glu + bp + bmi + age, You just need to place the known truth and your predicted probabilities (or other numerical predictor variable) into a data frame and then hand to the geom. For ggplot2, the package plotROC provides generic ROC plotting capabilities that work with any fitted model. Since you don't provide a reproducible example, I'll use a different dataset and model. And I don't have previous experience with R. Sorry if I am asking a really stupid question, but I don't have much time left for my project to finish. I would like to try code below by Claus Wilke however I got confused as I only have my main data (df) and my model (fit). (fit<-train(Attrition~.,data=df,method="glm",family="binomial",trControl=tc)) #train model, predict Attrition with all other variables Tc <- trainControl("cv",10,savePred=T) #create folds UPDATE: After viewing k-fold cross validation - how to get the prediction automatically? I changed my code to below: library("caret", lib.loc="~/R/win-library/3.4") I would be really grateful if you can help me with this. My problem is that, pred does not show up as a prediction, instead it shows as a data table. Pred = predict (modelfit, newdata=dt3Test)ĬonfusionMatrix(data=pred, dt3Test$Attrition) Modelfit <- train (Attrition~., data=dt3, method="glm", family="binomial", trControl=ctrl) I have created a logistic regression model with k-fold cross validation.ĭt3Training - training split made from main datasetĭt3Test - test split made from main datasetīelow is the code that used for logistic regression: ctrl<- trainControl (method="repeatedcv", number = 10, repeats =5, savePredictions="TRUE" I would like to know how can I draw a ROC plot with R. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
